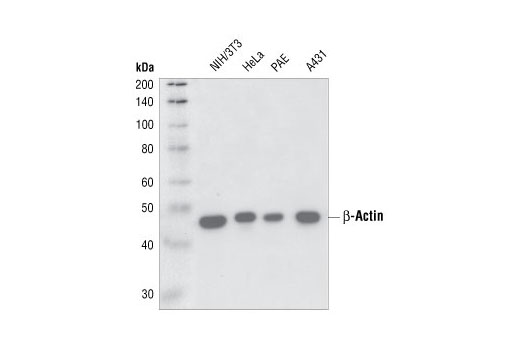 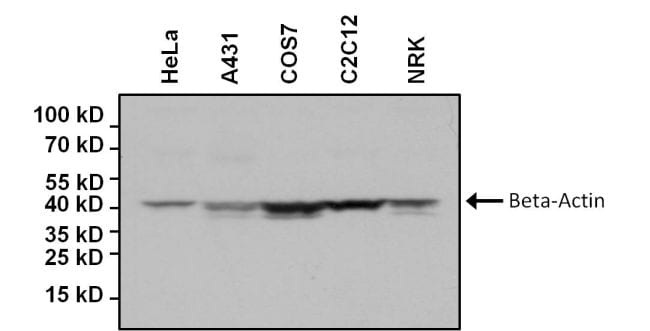
Protein heterogeneity, resulting in broad bands on the WB and uncertainties in the theoretical intact molecular weight of the spike protein, is assumed to be due to glycosylation. However, the identity of the bands identified by WB are not sufficiently justified and further clarification is requested.įurther in the document they say (emphasis mine):Īdditional data for the active substance are to be provided to confirm the identities of the observed Western Blot (WB) bands obtained by the in vitro expression assay. The expressed protein size after in-vitro expression of BNT162b2 active substance was determined and the results demonstrate comparability between batches. Unfortunately, I couldn’t find any images of western blots performed by the vaccine manufacturers, though we know they were performed.Īccording to an “ Assessment report ” for the Comirnaty (Pfizer-BioNTech) vaccine there are references to western blots (emphasis mine): 5 The sizes of the proteins we get from the vaccines don’t quite match with what’s predicted The window of opportunity for doing this is closing, since we'd ideally do this with people who never had COVID. It would be great if some researchers would follow this up and sequence spike from more vaccinated people to answer this and get a sense for how prevalent it is. Would we also find mutant peptides in some vaccinated people who didn’t have long COVID-like symptoms? Why would we have translation errors in S1 but not S2? Just a coincidence? Or something to do with the order in which the mRNA transcripts are read?Īnother possibility is that the vaccine mRNA had errors, and something about the vaccine mRNA manufacturing process caused more errors in the part that encodes for S1 compared to S2.Īll of these patients had long COVID-like symptoms, which presumably were from the vaccines. īut the cells of these patients had S2 subunits as well, and those did not contain mutant sequences. After all, the mRNA of the COVID vaccines had all their uridines replaced with N1-Methylpseudouridine, and there’s literature that suggests this can lead to errors in translation (see here and here). Perhaps this was due to errors that occurred while mRNA was translated into amino acids. The bottom shows the mutant (unexpected) S1 subunit peptides. The top shows expected S1 subunit peptides that were sequenced. “Peptide” just means a string of amino acid sequences. In other words, the S1 in these patients had amino acid sequences that were not intended by the vaccine manufacturers.įrom Fig 5B in Patterson et al. We know what the amino acid sequence of spike protein from the vaccines should be, because we know what the COVID vaccines are supposed to encode for.Īll six of the patients had mutant spike sequences, specifically in the S1 subunit of the spike (the other subunit is S2). 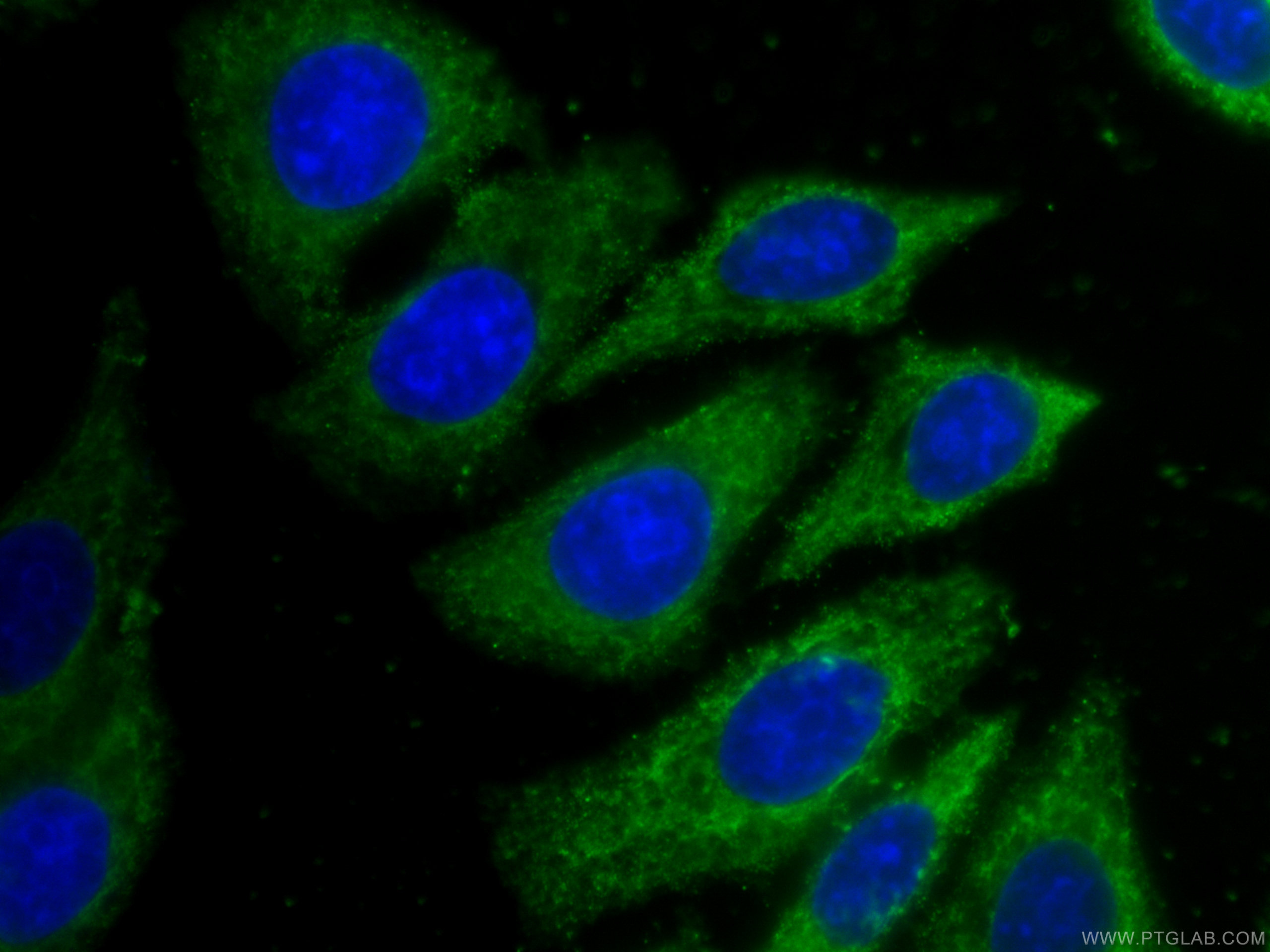
Remember: all proteins are made up of a string of amino acids. Of those patients, they chose six patients to isolate spike protein from their cells and determine its amino acid sequence. They found that at least 13 of the 14 patients had spike in their cells. They took 14 of these individuals and screened them for the presence of spike protein in a subset of their white blood cells. This paper looked at people who had not been previously infected with SARS-CoV-2, 2 had gotten vaccinated, 3 and had ongoing long COVID-like symptoms for more than 4 weeks following vaccination. In July 2022, a paper came out that should have gotten a lot more attention than it did: SARS-CoV-2 S1 Protein Persistence in SARS-CoV-2 Negative Post-Vaccination Individuals with Long COVID/ PASC-Like Symptoms
0 Comments
Leave a Reply. |
AuthorWrite something about yourself. No need to be fancy, just an overview. ArchivesCategories |
 RSS Feed
RSS Feed
